Artifact Removal Strategies#
GastroPy provides three composable time-domain preprocessing functions
that remove artifacts from the raw EGG signal before bandpass filtering,
plus a named pipeline variant in egg_clean that bundles them together.
Function |
Method |
Reference |
|---|---|---|
|
Sliding-window median; replaces spikes with local median |
Davies & Gather (1993); Dalmaijer (2025) |
|
Global MAD outlier replacement; faster, less adaptive |
Dalmaijer (2025) |
|
LMMSE Wiener filter for movement noise |
Gharibans et al. (2018) |
|
Hampel → LMMSE → IIR Butterworth pipeline |
Dalmaijer (2025) |
import matplotlib.pyplot as plt
import numpy as np
import gastropy as gp
plt.rcParams["figure.dpi"] = 100
plt.rcParams["figure.facecolor"] = "white"
# Synthetic EGG: gastric sine + pink noise + injected spike
rng = np.random.default_rng(42)
sfreq = 10.0
t = np.arange(0, 300, 1.0 / sfreq)
gastric = np.sin(2 * np.pi * 0.05 * t) # 3 cpm signal
noise = 0.3 * rng.standard_normal(len(t)) # background noise
movement = 0.5 * np.sin(2 * np.pi * 0.008 * t) # slow drift / movement
signal = gastric + noise + movement
# Inject two large spike artifacts
signal[300] = 8.0
signal[1800] = -7.5
print(f"Signal length : {len(signal)} samples ({len(signal)/sfreq:.0f} s)")
print(f"Signal range : [{signal.min():.2f}, {signal.max():.2f}]")
Signal length : 3000 samples (300 s)
Signal range : [-7.50, 8.00]
Hampel Filter — Spike Removal#
hampel_filter slides a window of length 2k+1 across the signal.
Any sample deviating from the local median by more than
n_sigma × 1.4826 × MAD is replaced by that median.
hampel_cleaned = gp.hampel_filter(signal, k=5, n_sigma=3.0)
fig, axes = plt.subplots(2, 1, figsize=(10, 5), sharex=True)
axes[0].plot(t, signal, lw=0.8, color="#d62728", label="Raw (with spikes)")
axes[0].set_ylabel("Amplitude (µV)")
axes[0].legend(loc="upper right")
axes[1].plot(t, hampel_cleaned, lw=0.8, color="#1f77b4", label="Hampel filtered")
axes[1].set_ylabel("Amplitude (µV)")
axes[1].set_xlabel("Time (s)")
axes[1].legend(loc="upper right")
fig.suptitle("Hampel Filter — Spike Removal")
plt.tight_layout()
plt.show()
print(f"Max absolute value before: {np.abs(signal).max():.2f}")
print(f"Max absolute value after : {np.abs(hampel_cleaned).max():.2f}")
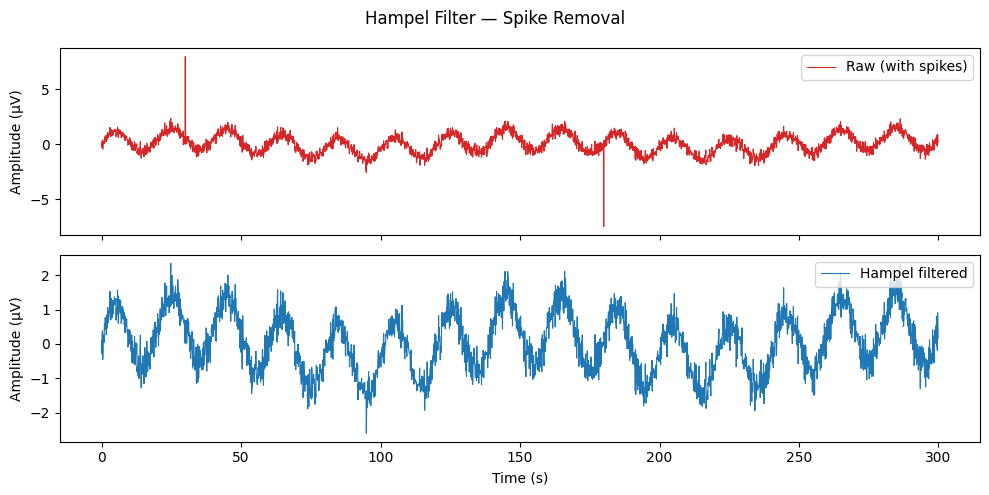
Max absolute value before: 8.00
Max absolute value after : 2.59
MAD Filter — Global Outlier Removal#
mad_filter computes a single global median and MAD across the entire
signal. Faster than hampel_filter but less adaptive to slow signal
drift. Suitable as a quick first pass on stationary recordings.
mad_cleaned = gp.mad_filter(signal, n_sigma=3.0)
fig, axes = plt.subplots(2, 1, figsize=(10, 5), sharex=True)
axes[0].plot(t, signal, lw=0.8, color="#d62728", label="Raw (with spikes)")
axes[0].set_ylabel("Amplitude (µV)")
axes[0].legend(loc="upper right")
axes[1].plot(t, mad_cleaned, lw=0.8, color="#2ca02c", label="MAD filtered")
axes[1].set_ylabel("Amplitude (µV)")
axes[1].set_xlabel("Time (s)")
axes[1].legend(loc="upper right")
fig.suptitle("MAD Filter — Global Outlier Removal")
plt.tight_layout()
plt.show()
print(f"Max absolute value after MAD filter: {np.abs(mad_cleaned).max():.2f}")
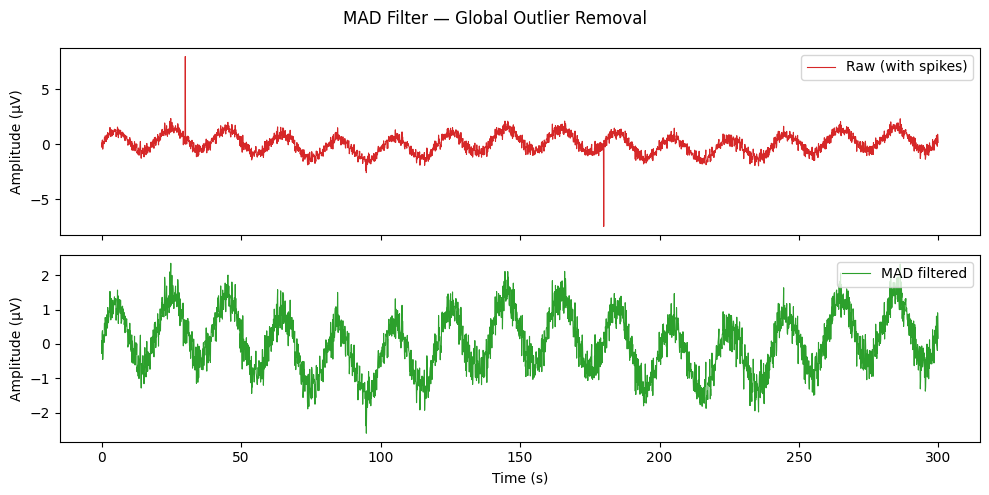
Max absolute value after MAD filter: 2.59
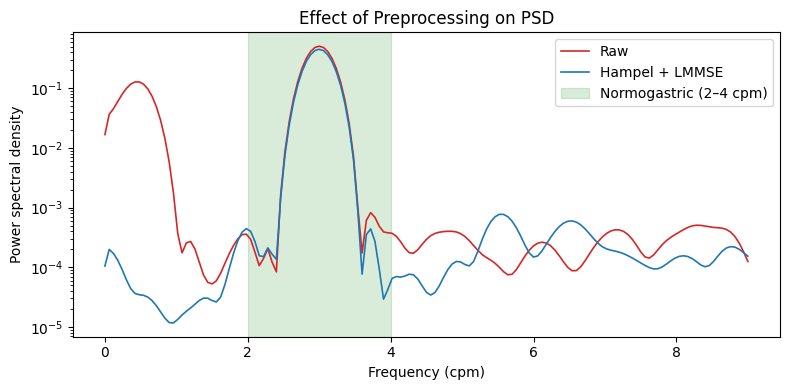
Movement Artifact Filter — LMMSE#
remove_movement_artifacts applies an LMMSE Wiener-filter that
estimates local variance in sliding windows and attenuates time segments
where variance exceeds the expected signal variance. Effective for
movement and drift artifacts (Gharibans et al., 2018).
# First remove spikes, then attenuate movement artifacts
spike_cleaned = gp.hampel_filter(signal, k=5)
movement_cleaned = gp.remove_movement_artifacts(spike_cleaned, sfreq=sfreq, freq=0.05)
# Compare PSDs
freqs_raw, psd_raw = gp.psd_welch(signal, sfreq, fmin=0.0, fmax=0.15, overlap=0.75)
freqs_clean, psd_clean = gp.psd_welch(movement_cleaned, sfreq, fmin=0.0, fmax=0.15, overlap=0.75)
fig, ax = plt.subplots(figsize=(8, 4))
ax.semilogy(freqs_raw * 60, psd_raw, color="#d62728", lw=1.2, label="Raw")
ax.semilogy(freqs_clean * 60, psd_clean, color="#1f77b4", lw=1.2, label="Hampel + LMMSE")
ax.axvspan(2, 4, alpha=0.15, color="green", label="Normogastric (2–4 cpm)")
ax.set_xlabel("Frequency (cpm)")
ax.set_ylabel("Power spectral density")
ax.set_title("Effect of Preprocessing on PSD")
ax.legend()
plt.tight_layout()
plt.show()
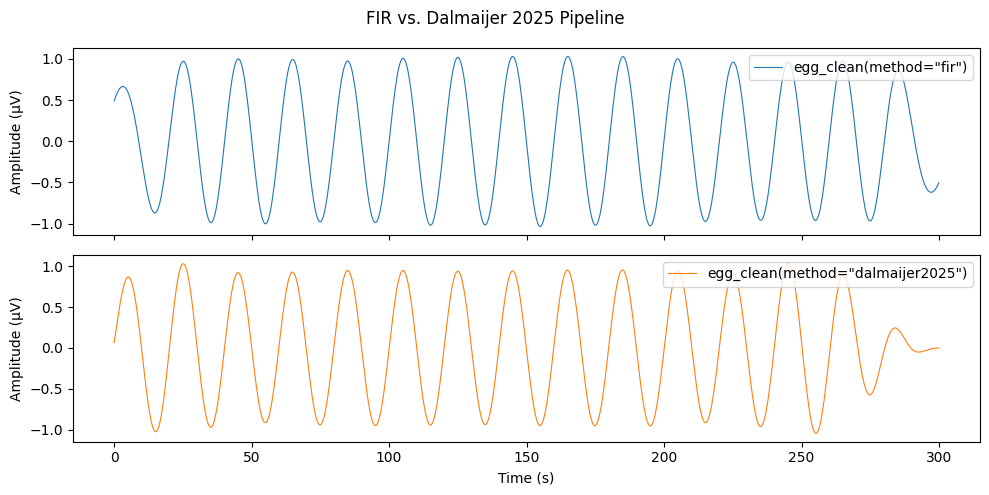
Dalmaijer 2025 — Bundled Pipeline#
egg_clean(method="dalmaijer2025") runs Hampel → LMMSE → IIR
Butterworth as a single named pipeline. Compare with the default FIR
bandpass on the same noisy signal.
cleaned_fir, info_fir = gp.egg_clean(signal, sfreq=sfreq, method="fir")
cleaned_d25, info_d25 = gp.egg_clean(signal, sfreq=sfreq, method="dalmaijer2025")
print("FIR pipeline:")
print(f" filter_method : {info_fir['filter_method']}")
print(f" numtaps : {info_fir['fir_numtaps']}")
print("\nDalmaijer 2025 pipeline:")
print(f" cleaning_method : {info_d25['cleaning_method']}")
print(f" hampel_k : {info_d25['hampel_k']}")
print(f" movement_window : {info_d25['movement_window']} cycles")
print(f" filter_method : {info_d25['filter_method']}")
fig, axes = plt.subplots(2, 1, figsize=(10, 5), sharex=True)
axes[0].plot(t, cleaned_fir, lw=0.8, color="#1f77b4", label='egg_clean(method="fir")')
axes[0].set_ylabel("Amplitude (µV)")
axes[0].legend(loc="upper right")
axes[1].plot(t, cleaned_d25, lw=0.8, color="#ff7f0e", label='egg_clean(method="dalmaijer2025")')
axes[1].set_ylabel("Amplitude (µV)")
axes[1].set_xlabel("Time (s)")
axes[1].legend(loc="upper right")
fig.suptitle("FIR vs. Dalmaijer 2025 Pipeline")
plt.tight_layout()
plt.show()
FIR pipeline:
filter_method : fir
numtaps : 501
Dalmaijer 2025 pipeline:
cleaning_method : dalmaijer2025
hampel_k : 3
movement_window : 1.0 cycles
filter_method : iir_butter
See also: Bandpass Filtering, One-Call EGG Pipeline, Multi-Channel Processing, Artifact Detection
